 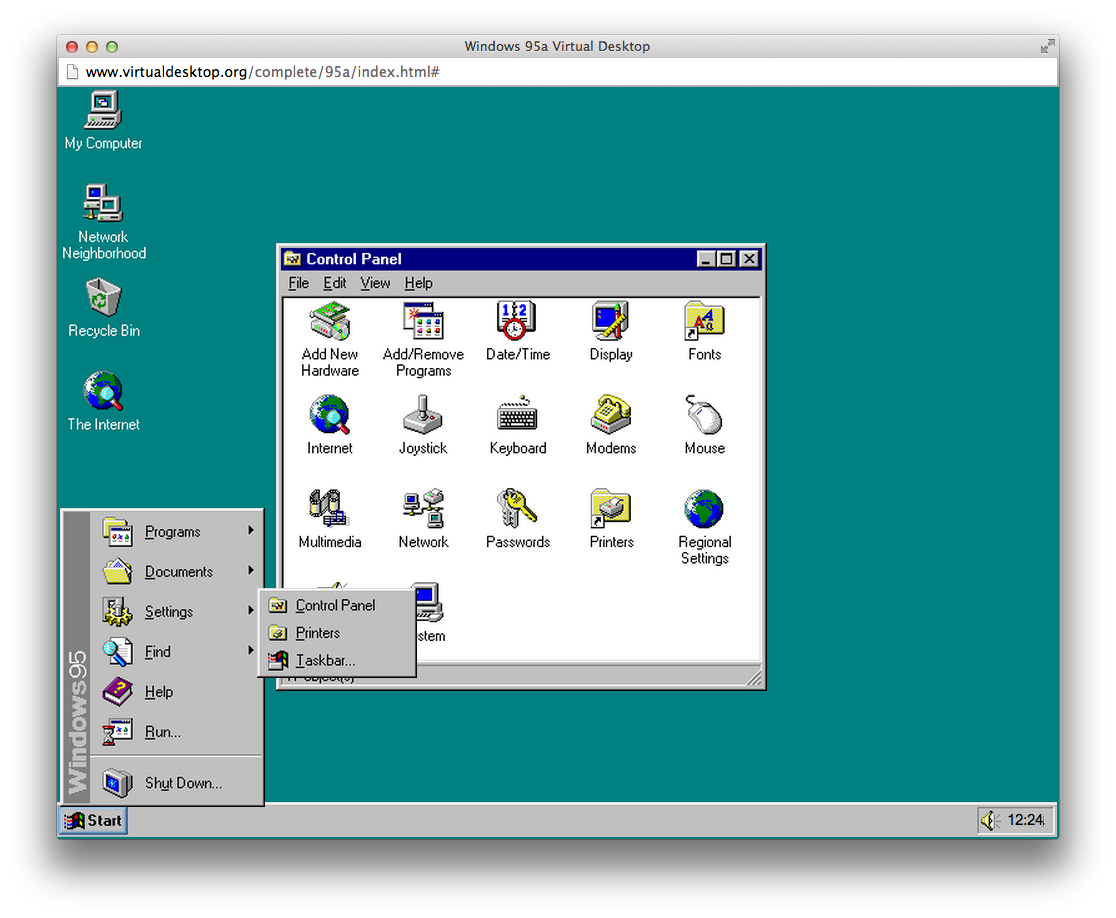 
Genome Workbench GUI before the editing package is enabled. Log out and log back in to enable xauth, see notes for connecting from Windows or Mac.logout and log back in to enable xauth, see notes for connecting from Windows or Mac.“log out and log back in…” Installing on CentOs 7 Be aware of the accidental line-breaks – each step should constitute exactly one command – with the obvious exception of the instructions to the user, e.g. Some AWS images may not have even these tools available - CoreOs or AtomicHost versions of Fedora for example.Īfter launching and logging in to your instance via ssh, please issue each command from the command line exactly as written. The version of Linux running on an AWS instance needs to have some minimal set of tools installed (yum for CentOs, dnf for Fedora, zypper for OpenSUSE, apt for Ubuntu and Debian) as well as have sshd running. You may select an instance with more or less RAM available, depending on your computational needs. We used t2.large instance (8Gb RAM) for testing. If your Linux host or AWS instance is running on an ARM CPU, you will have to build the software from source. At the moment, the only supported architecture is x86_64. As updated versions are made available, you may have to adjust the download URLs in the steps below. You can always find and download the latest release. The steps below assume Genome Workbench version 3.0.0. Please note that configuration and launching of instances in AWS EC2 as well as the detailed tutorial on Genome Workbench usage for specific tasks is beyond the scope of this document. Please see the Connecting to Your Linux Instance from Windows Using PuTTY notes if you are not already conversant with ssh. You also need to be familiar with the procedure to connect to the EC2 instances using ssh. See “ connecting from a Windows PC” or “ connecting from a Mac” directions to connect to the Linux host from Windows or Mac and display the Genome Workbench GUI. This tutorial may also be useful if you are trying to install Genome Workbench on a Linux host running elsewhere. This guide gives step by step instructions on how to install Genome Workbench on Linux hosts running in the Amazon EC2 Cloud. A more cost effective approach would be to bring the software to the cloud instead of copying the data in the other direction. It may frequently be a non-trivial expense to move such amounts of data between the cloud storage and a desktop computer. Genome data sets can reach tens and hundreds of gigabytes in size. If your data processing pipeline already exists on the cloud it might be faster and more convenient to add Genome Workbench to your toolbox there instead of installing it on your desktop PC or Mac. Why run Genome Workbench on the cloud? In addition to the usual benefits of being able to connect to the software from anywhere without the need to maintain your own server there are multiple other reasons. Running Genome Workbench over X Window System
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
